Filtering
VarLens provides a layered filtering system — from one-click presets to a structured query language — so you can narrow down variants at whatever level of detail you need.
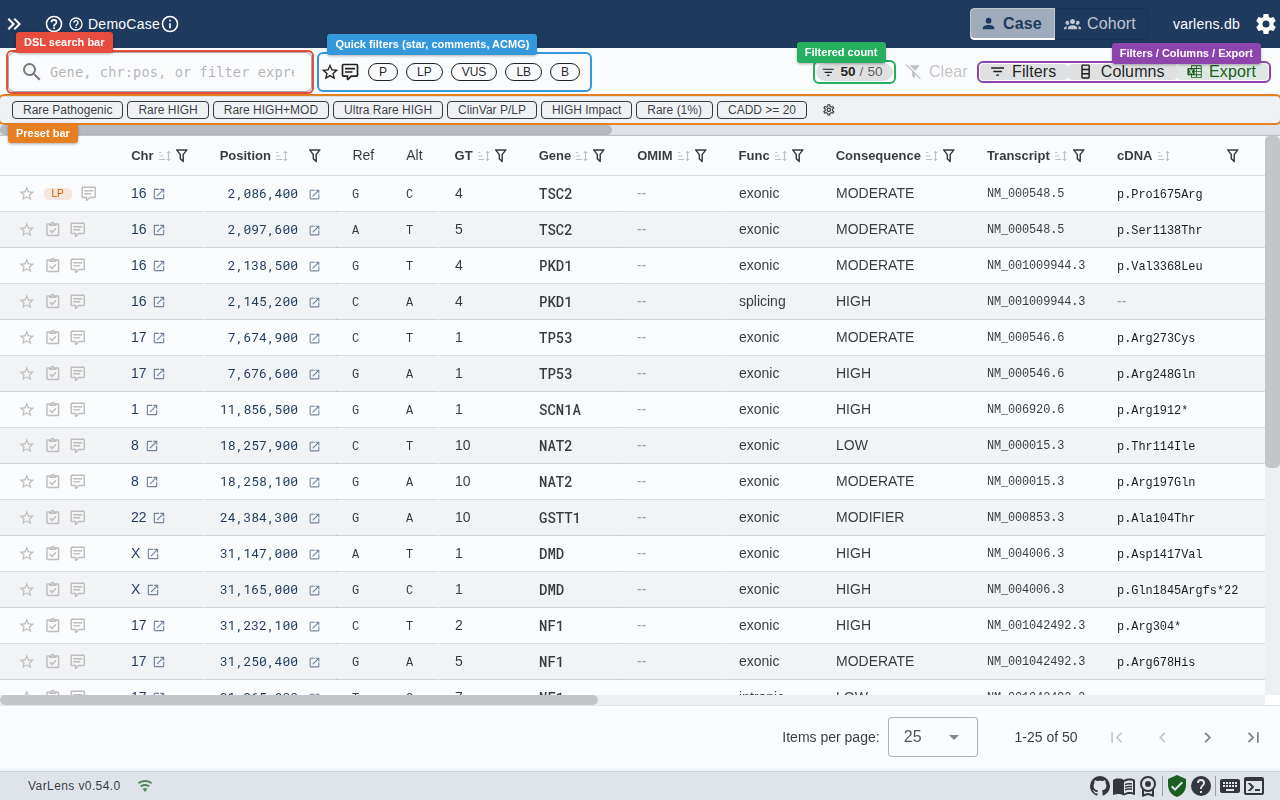
Quick Filters (Toolbar)
The toolbar above the variant table provides instant-access filters:
| Control | What it does |
|---|---|
| Search bar | Free-text search (gene, position, HGVS) or DSL expressions |
| Case / All | Scope annotation filters to current case or all cases |
| Star / Comment | Show only starred or commented variants |
| ACMG chips | Filter by classification (P, LP, VUS, LB, B) |
| Tags | Filter by assigned tags |
The result count chip shows how many variants pass your filters (e.g., 12 / 245). It pulses briefly when the count changes.
Active Filter Bar
When filters are active, a bar appears below the toolbar showing each filter as a removable chip:
- Click the × on any chip to remove that filter
- Click Clear all to reset everything
Filter Drawer
Click Filters or press Ctrl+Shift+F to open the drawer. Filters are grouped into sections:
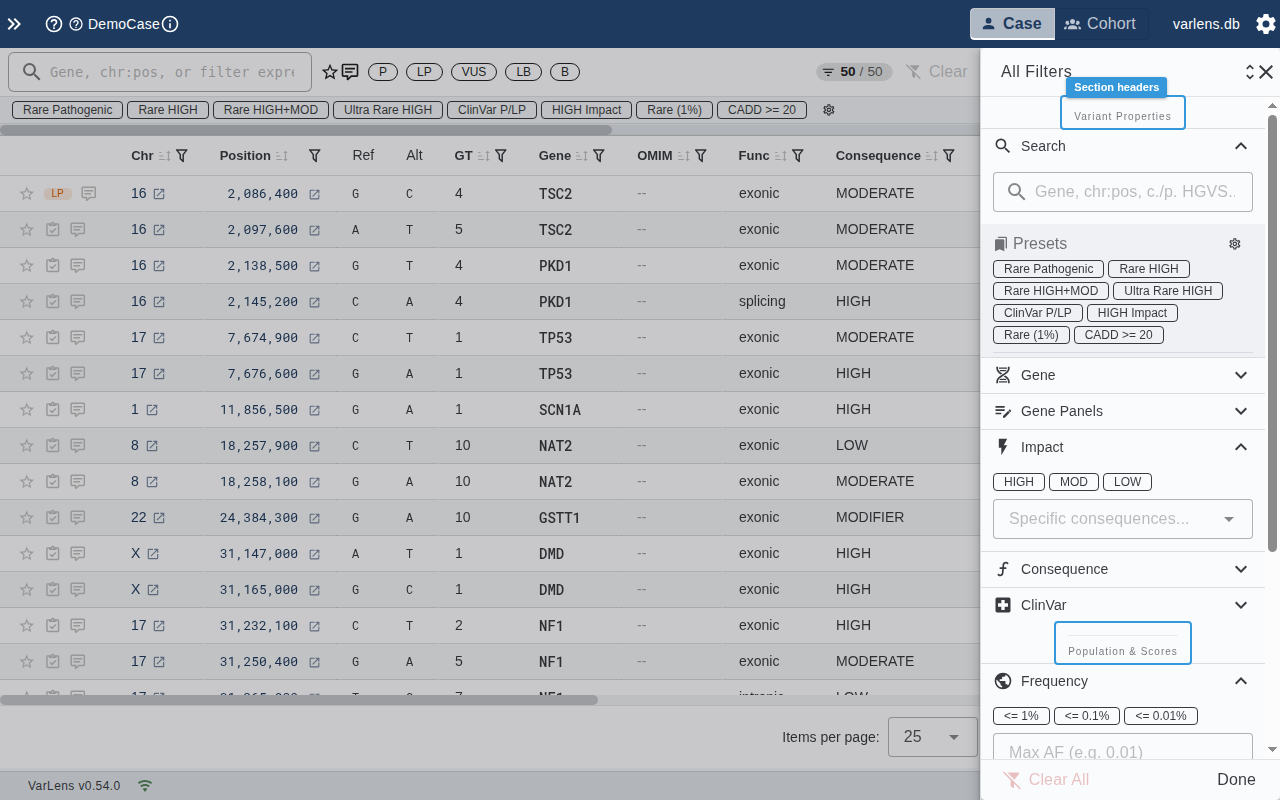
Variant Properties
- Search — Full-text search across all text fields
- Gene — Autocomplete by gene symbol
- Impact — Toggle HIGH / MODERATE / LOW impact chips
- Consequence — Grouped multi-select (Truncating, Missense, Splice, etc.)
- ClinVar — Grouped multi-select (Pathogenic, VUS, Benign, etc.)
Population & Scores
- Frequency — gnomAD allele frequency threshold (presets: ≤ 1%, ≤ 0.1%, ≤ 0.01%) with custom input
- CADD — Minimum CADD Phred score (presets: ≥ 10, ≥ 15, ≥ 20, ≥ 25) with custom input
Numeric filters are NULL-inclusive by default: variants without annotation data (e.g., novel variants with no gnomAD entry) pass through frequency and CADD filters.
Annotations
- Tags — Filter by assigned tags
- Star / Comment — Toggle starred or commented variants
- ACMG — Filter by ACMG classification
Collapsed Previews
When a filter panel is collapsed, its current value appears on the right (e.g., "≤ 1.00%" next to Frequency). This lets you see all active filters at a glance.
Per-Column Filters
Each column header has a filter icon. Click it to open a type-aware filter popup:
Numeric Columns (CADD, gnomAD AF, Quality)
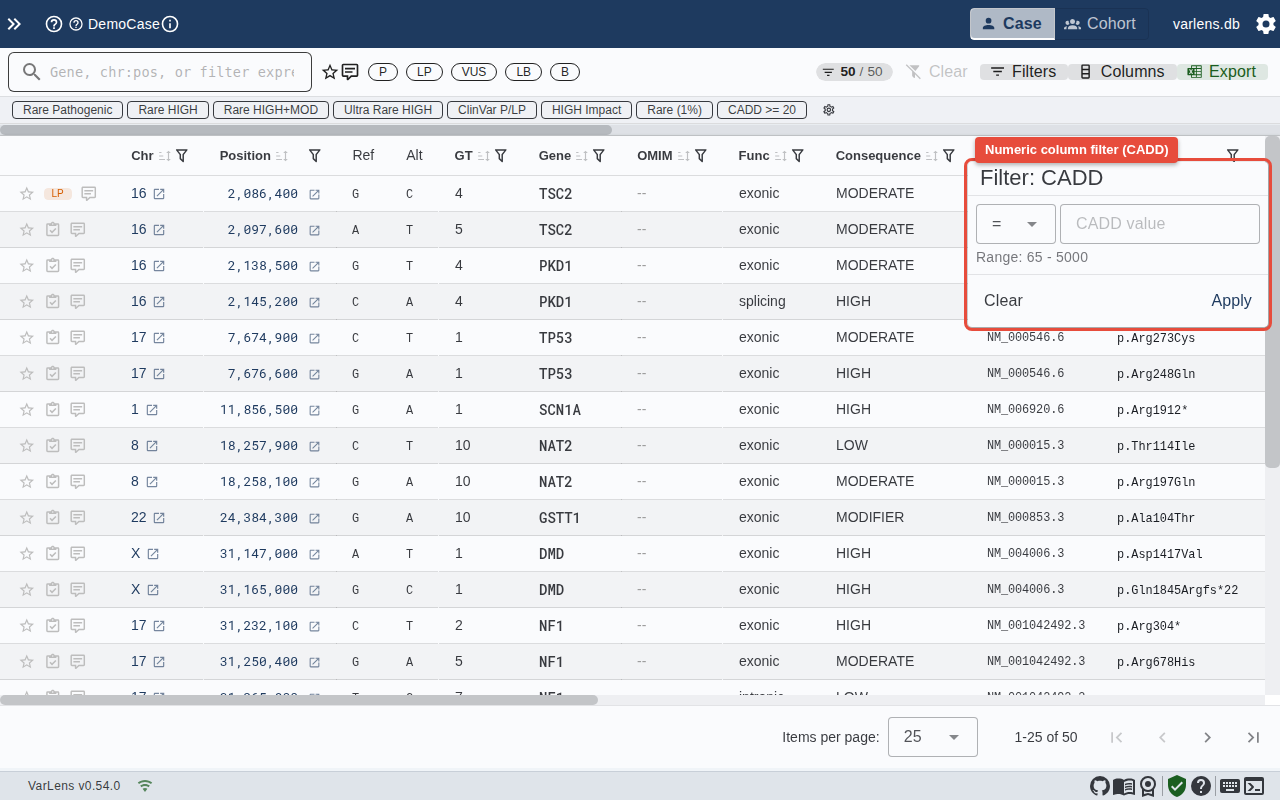
- Choose an operator (
<,>,<=,>=,=,!=) - Enter a value or click a preset
- Toggle "Include missing values" for NULL-inclusive behavior
- The data range in the current case is shown at the bottom
Categorical Columns (Consequence, ClinVar, Function)
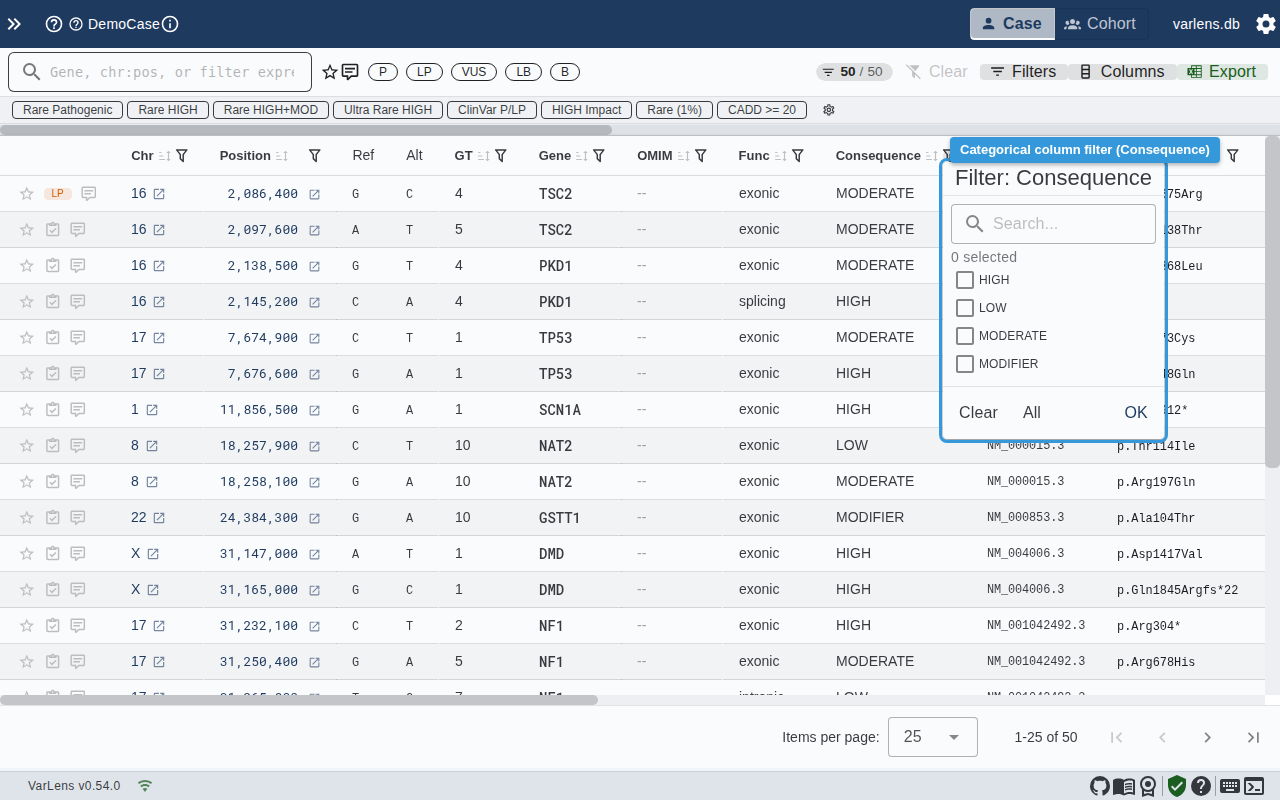
- Search within available values
- Check/uncheck individual values (with counts)
- Use "Select All" / "Clear" for bulk operations
Text Columns (Gene, Transcript, cDNA)
- Choose a match mode: Contains, Equals, Starts with, Ends with
- Type a search term — matching values are previewed
DSL Search Bar
For power users, the search bar supports a structured filter language:
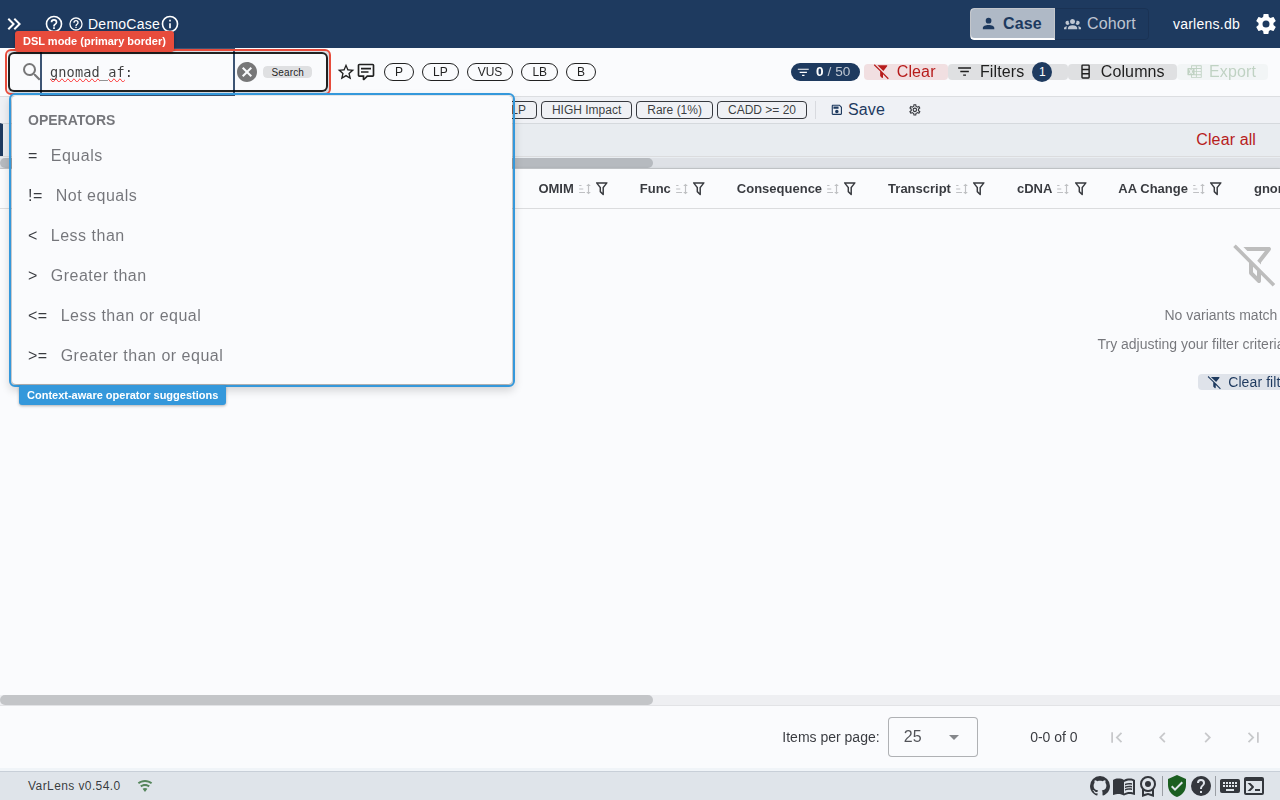
Syntax
column:operator:valueExamples:
| Expression | Meaning |
|---|---|
gnomad_af:<:0.01 | AF less than 1% |
cadd:>=:20 | CADD at least 20 |
gene:=:BRCA1 | Exact gene match |
consequence:~:missense | Consequence contains "missense" |
Operators
| Operator | Name | Column types |
|---|---|---|
= | Equals | All |
!= | Not equals | All |
< > <= >= | Comparisons | Numeric |
~ | Contains | Text, Categorical |
Combining Filters
Use AND or OR between expressions:
gnomad_af:<:0.01 AND cadd:>=:20Use parentheses to group OR conditions:
(gene:=:BRCA1 OR gene:=:TP53) AND gnomad_af:<:0.01WARNING
Mixing AND and OR without parentheses is not allowed — VarLens will ask you to add parentheses to clarify your intent.
Autocomplete
The search bar offers context-aware suggestions as you type:
- Column names — type a few letters to see matching columns
- Operators — after
column:, valid operators for that column type appear - Values — after
column:op:, common values are suggested (e.g., AF thresholds) - Combinators — after a complete expression,
AND/ORare suggested
Preset References
Type @ followed by a preset name to apply a saved preset directly from the search bar:
@rare-pathogenicPlain Text Search
If your input doesn't contain colons, it works as a regular full-text search:
BRCA1— search by gene symbolBRCA1 AND pathogenic— boolean operatorsc.5123C>T— HGVS cDNA notationp.Ala1708Glu— HGVS protein notation
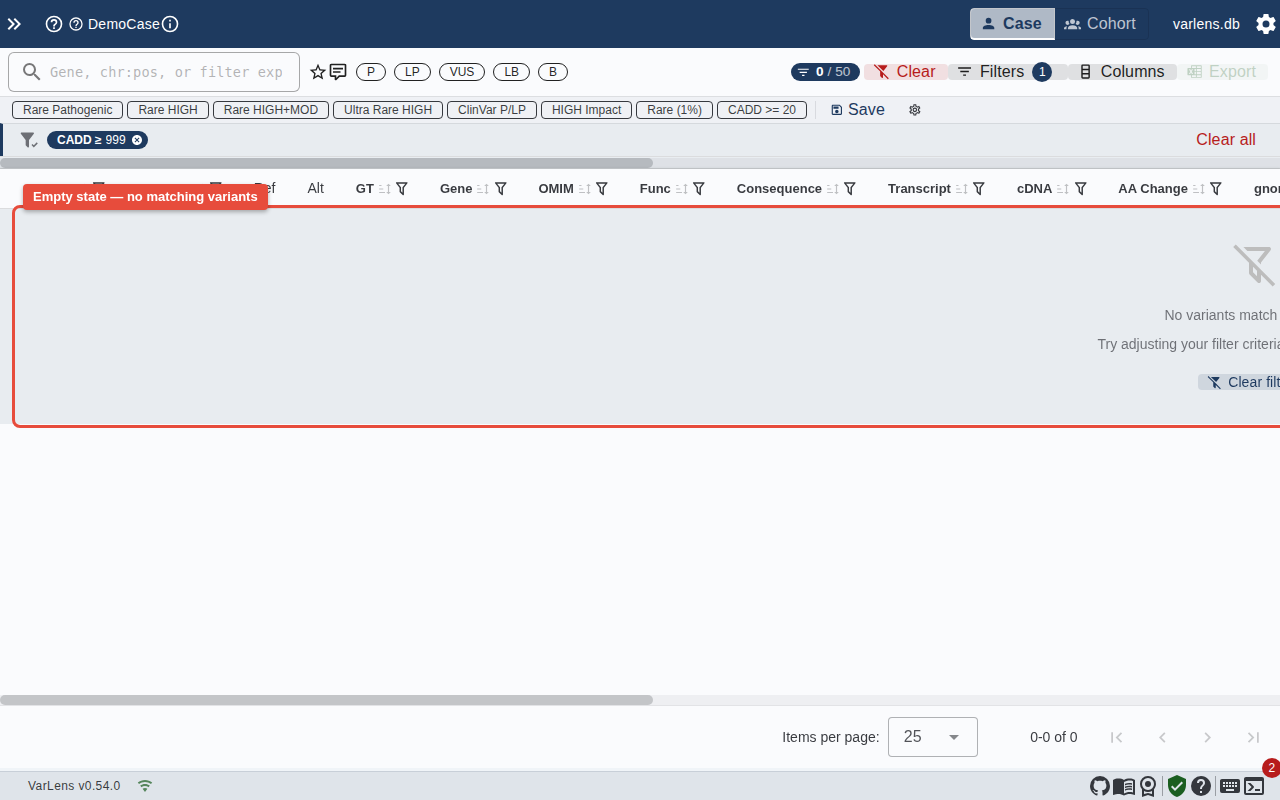
Empty State
When filters produce no matching variants, VarLens shows a clear message with a button to reset all filters:
See Also
- Filter Presets — save and reuse filter combinations
- Keyboard Shortcuts — filter-related shortcuts