Gene Panels
VarLens includes a gene panel system for region-based variant filtering. Create panels from gene lists, import from PanelApp, or generate interaction panels from StringDB — then apply them to filter variants by genomic coordinates.
Filter Sidebar
Select panels from the Gene Panels section in the filter drawer. Active panels restrict the variant table to genomic regions covered by the panel's genes (with configurable padding).
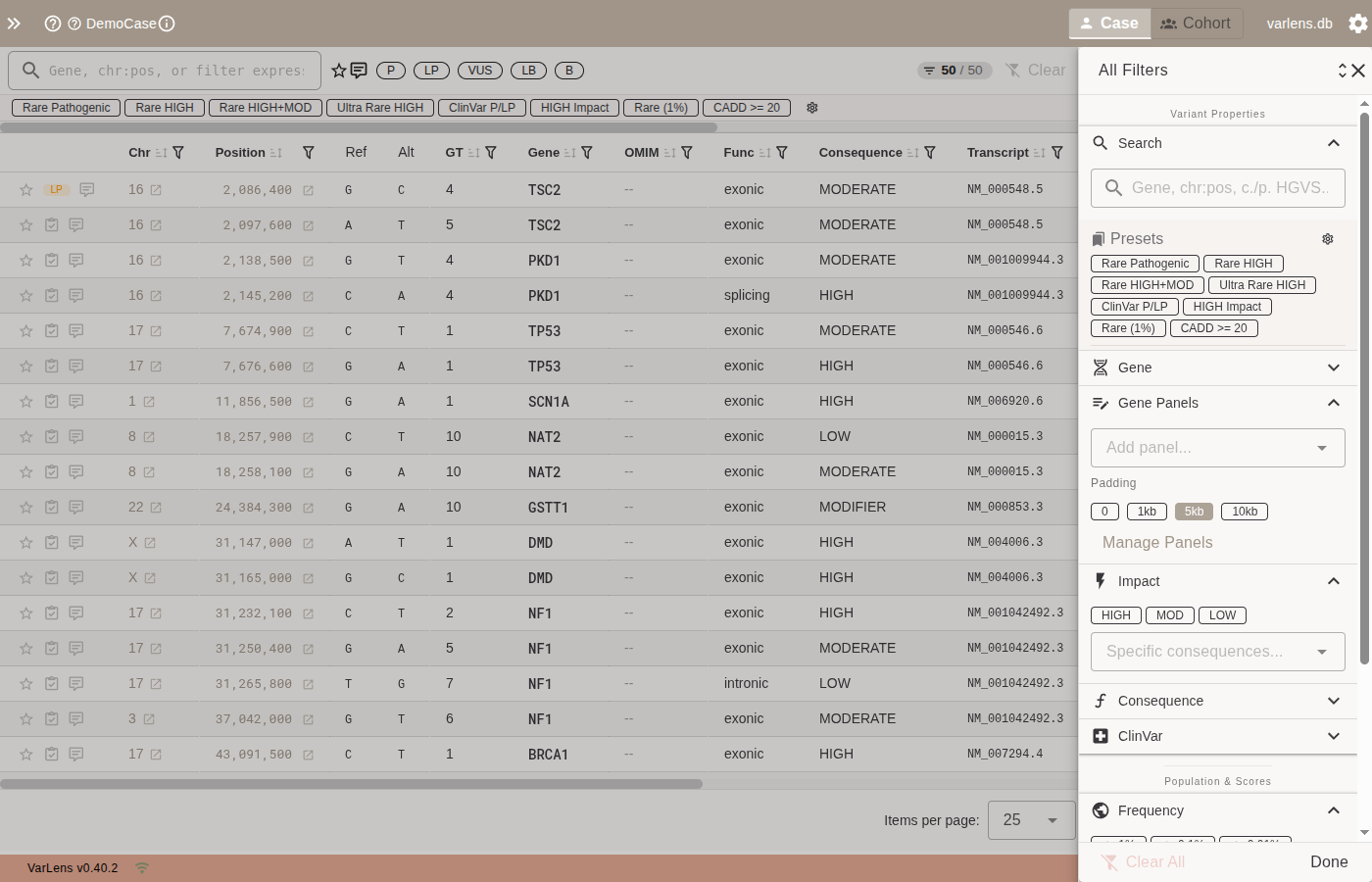
| Control | Description |
|---|---|
| Add panel | Dropdown to select a panel for filtering |
| Padding | Base pairs added around each gene (0, 1kb, 5kb, 10kb) |
| Manage Panels | Opens the panel management dialog |
Multiple panels can be active simultaneously — their regions are merged (union).
Panel Manager
Access via Settings > Gene Panels or Manage Panels in the filter drawer.
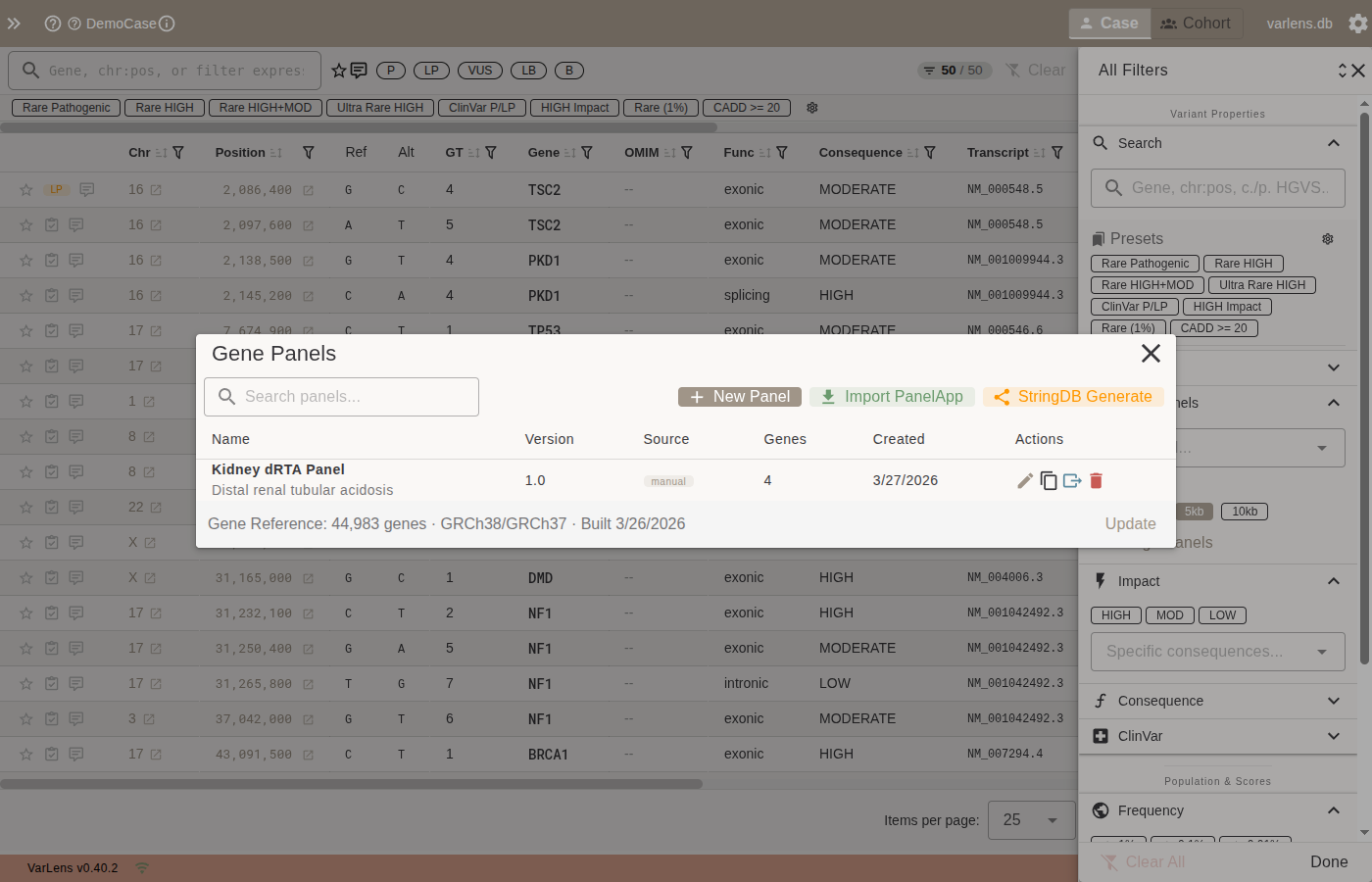
The manager shows all panels with name, version, source, gene count, and creation date. Actions per panel:
- Edit — modify name, version, description, or gene list
- Copy — duplicate the panel
- Export — save as BED file (choose assembly and padding)
- Delete — remove with confirmation
The footer shows the bundled gene reference database status (gene count, assemblies, build date).
Creating Panels
Click New Panel to open the editor. Enter a name, optional version, and description, then add genes.
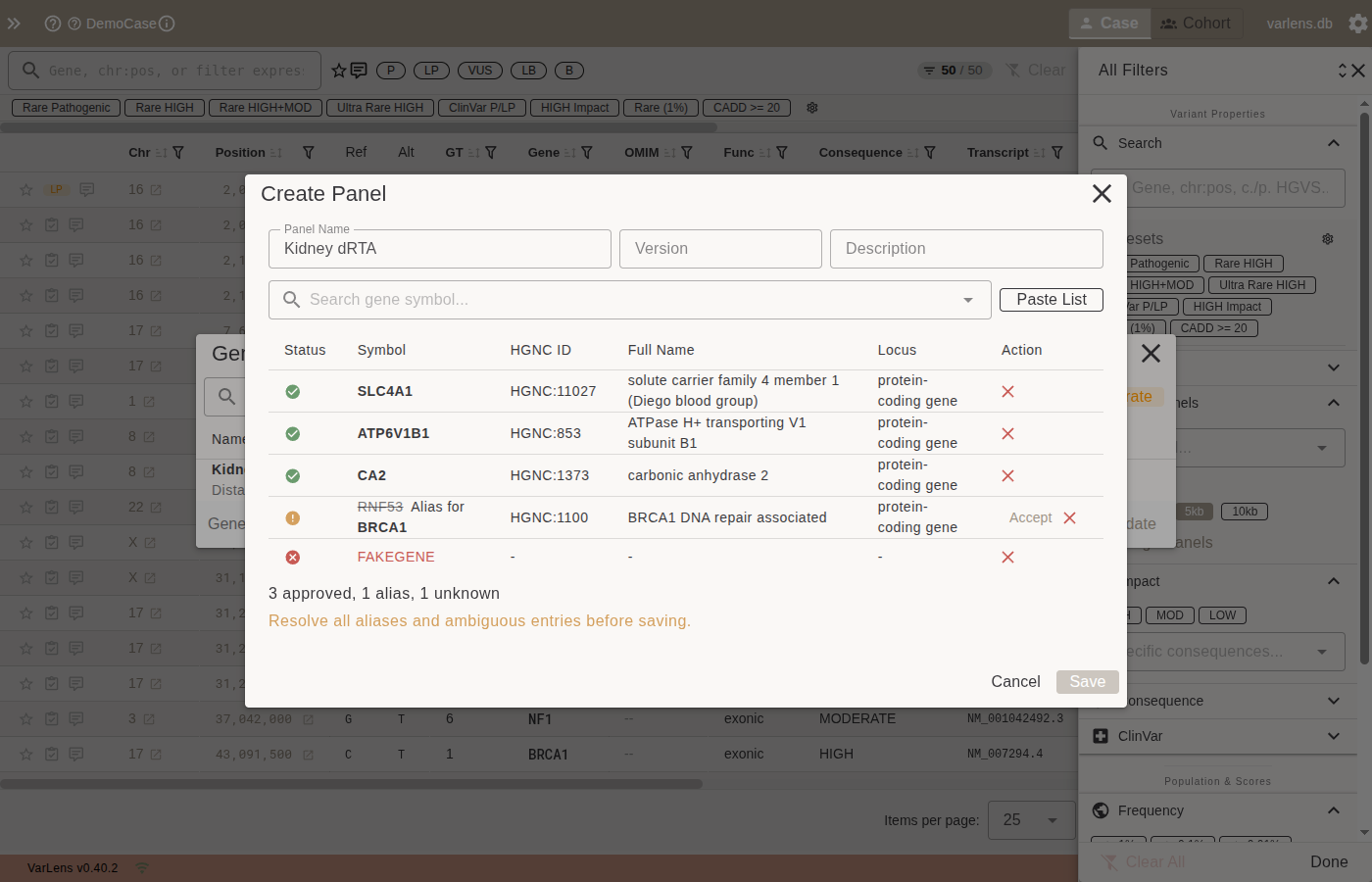
Adding Genes
Autocomplete search — type a gene symbol in the search field. Results show the HGNC-approved symbol, full name, and locus group. Alias matches display the matched alias.
Paste a list — click Paste List to bulk-add genes separated by newlines, commas, or semicolons. All symbols are validated against the bundled HGNC gene reference.
Validation States
| Status | Icon | Meaning |
|---|---|---|
| Approved | Green check | Current HGNC symbol, ready to save |
| Alias | Orange warning | Matched an alias or previous symbol — click Accept to resolve |
| Ambiguous | Orange warning | Alias maps to multiple genes — select the correct one |
| Unknown | Red X | No HGNC match — must be removed before saving |
Panels can only be saved when all genes are in the Approved state (strict validation).
Importing from PanelApp
Click Import PanelApp to search and import expert-curated disease gene panels from Genomics England PanelApp (UK) or PanelApp Australia.
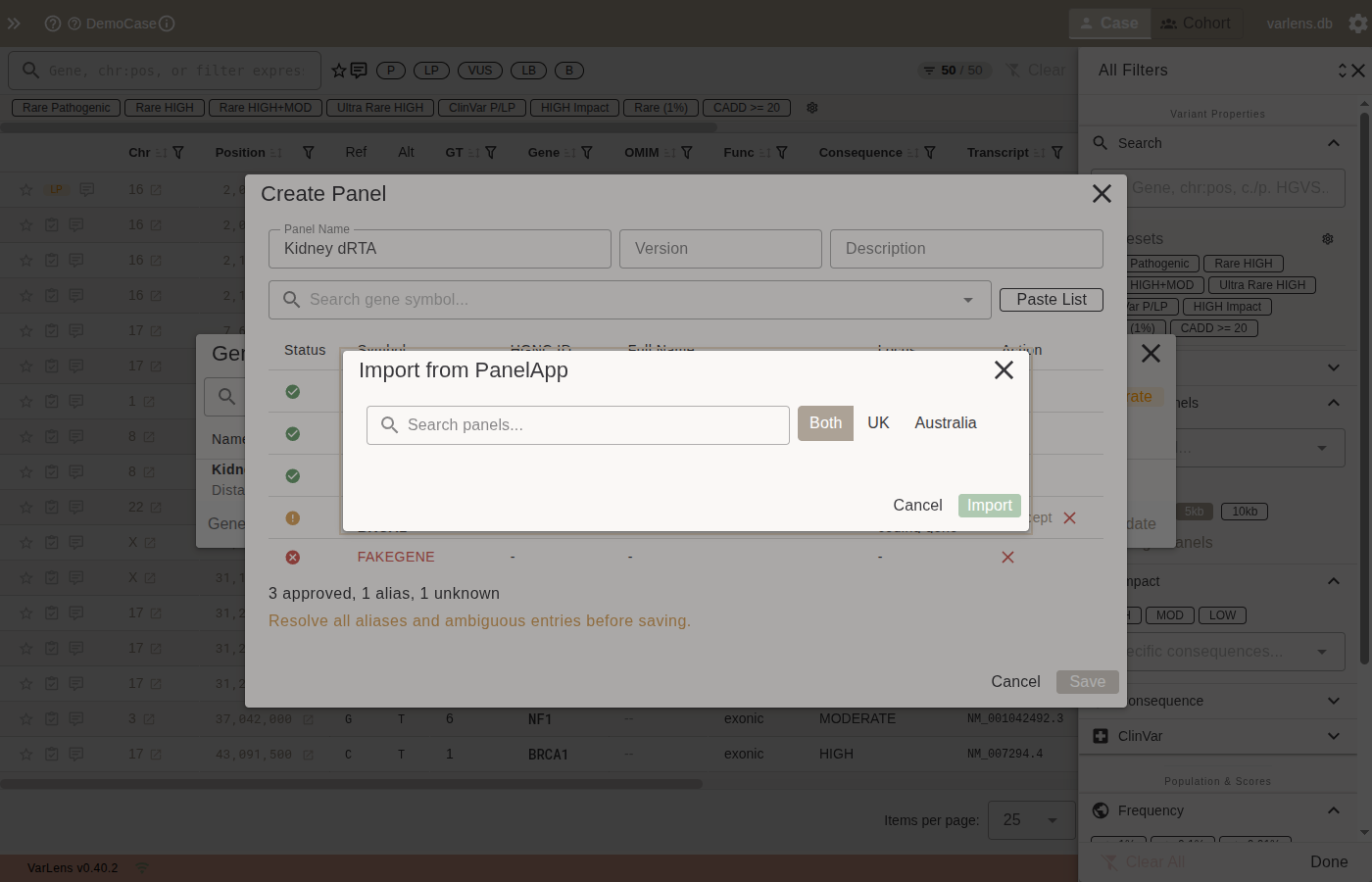
- Enter a keyword (e.g., "kidney") or panel ID
- Select region: UK, Australia, or Both
- Choose a panel from the results
- Set confidence level filter: Green only (diagnostic grade), Green + Amber, or All
- Click Import
Imported panels track their source, version, and confidence threshold.
Generating from StringDB
Click StringDB Generate to create interaction panels from protein-protein interaction data.
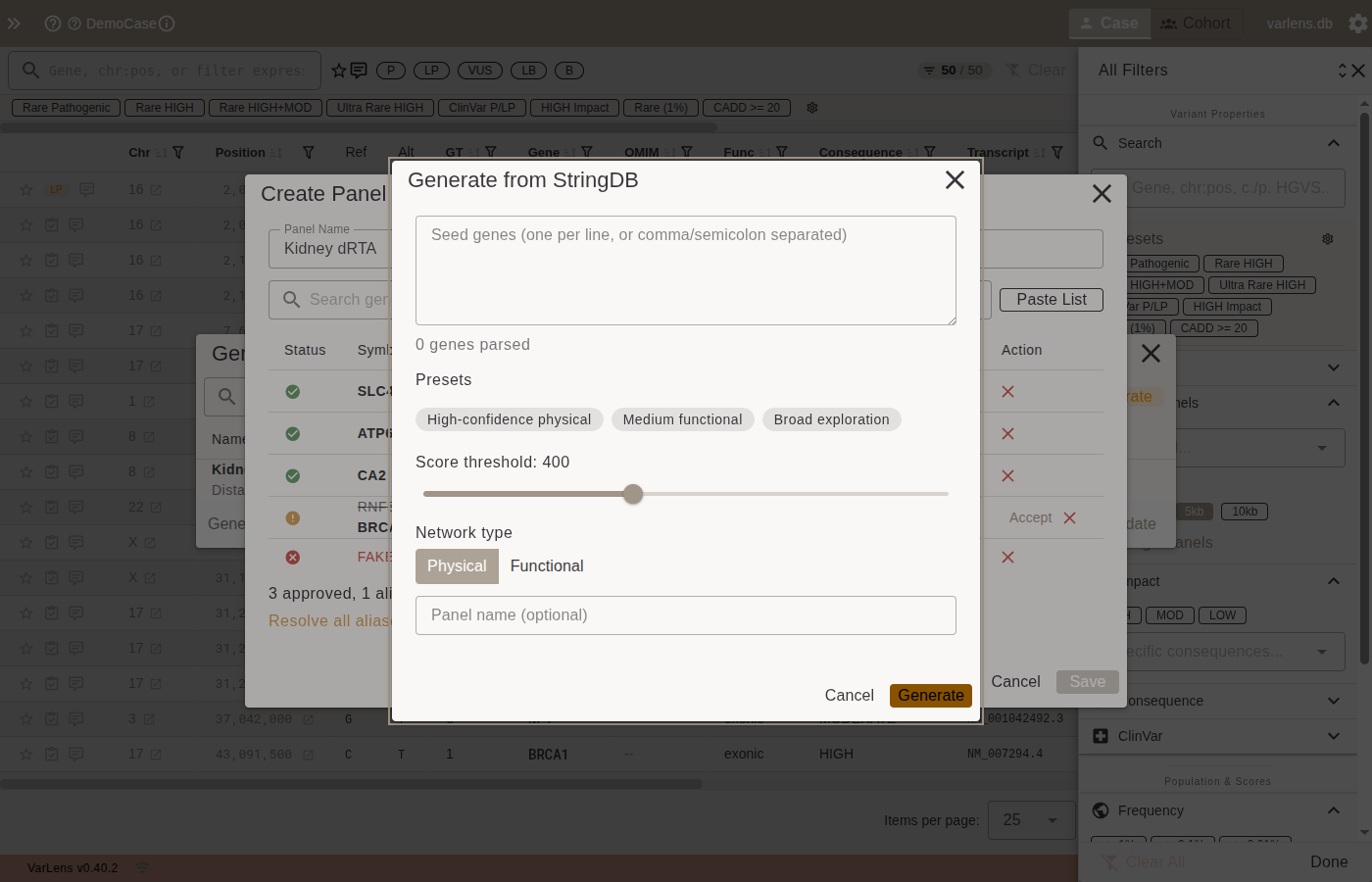
- Enter seed genes (paste or one per line)
- Select a preset or configure custom parameters:
- High-confidence physical — score >= 700, physical interactions only
- Medium functional — score >= 400, functional network
- Broad exploration — score >= 150, discovery mode
- Click Generate
The resulting panel includes the seed genes plus their validated interaction partners.
How Filtering Works
When a panel is active, VarLens:
- Looks up genomic coordinates for each gene in the panel using the bundled reference database
- Applies the selected padding (e.g., +/- 5kb) around each gene
- Merges overlapping intervals
- Filters variants to only those within the resulting genomic regions
The coordinates are matched to the case's genome build (GRCh37 or GRCh38). Panel filtering works in both case view and cohort view.
Gene Reference Database
VarLens ships with a bundled gene reference database containing:
- ~45,000 HGNC genes with symbols, aliases, and metadata
- Genomic coordinates for GRCh37 and GRCh38 (from Ensembl)
- FTS5 full-text search indexes for fast autocomplete
The database is extensible to future assemblies (e.g., T2T-CHM13).