Variant Table
The variant table is the primary view for analyzing variants in a case. It displays all imported variants in a sortable, scrollable data table with customizable columns.
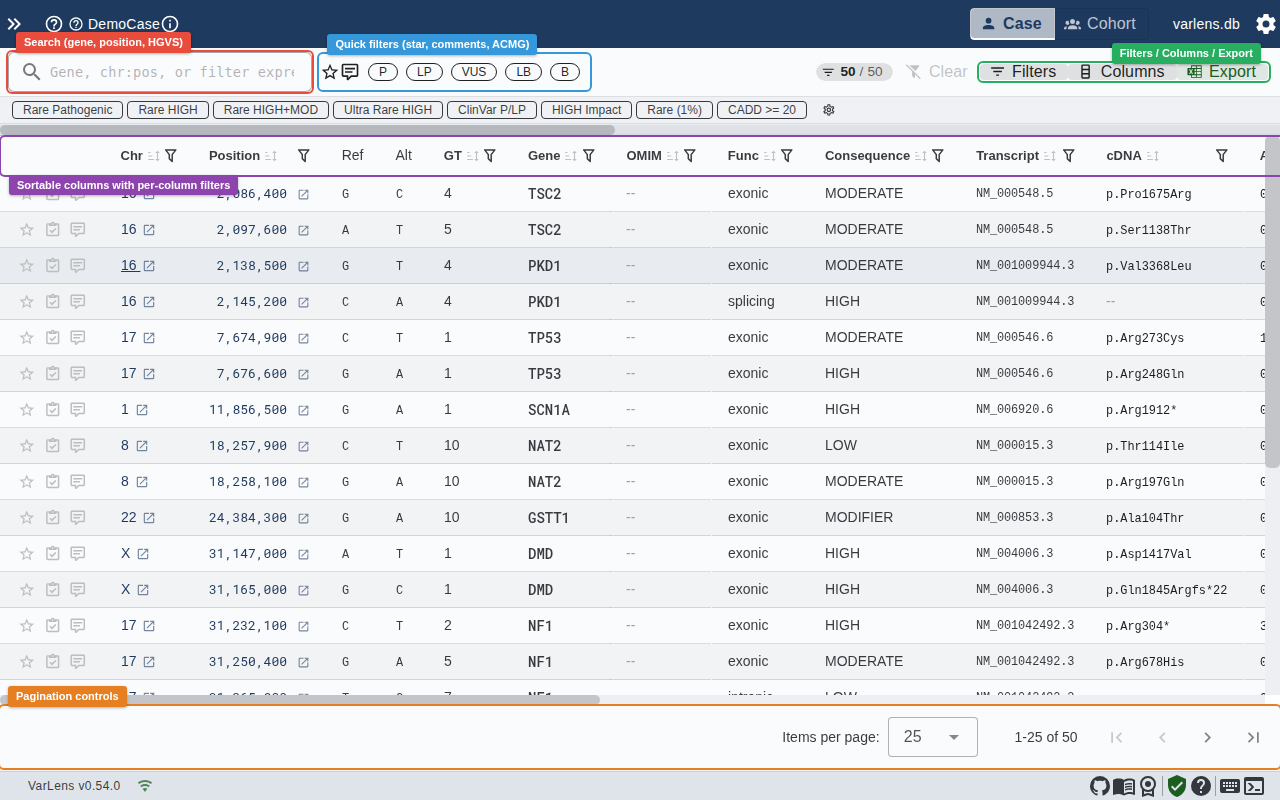
Columns
The table includes the following columns by default:
| Column | Description |
|---|---|
| Annotations | Star, ACMG classification, comments |
| Chr | Chromosome |
| Pos | Genomic position (formatted with separators) |
| Ref / Alt | Reference and alternate alleles |
| GT | Genotype (0/1 het, 1/1 hom) |
| Gene | Gene symbol |
| OMIM | OMIM disease number |
| Func | Functional class (exonic, splicing, intronic, etc.) |
| Consequence | Variant consequence with color coding |
| Transcript | Selected transcript ID |
| cDNA | HGVS coding DNA change |
| AA Change | HGVS protein change |
| gnomAD AF | Population allele frequency |
| CADD | CADD pathogenicity score |
| Qual | Variant call quality score |
| ClinVar | ClinVar clinical significance |
| HPO Sim | HPO similarity score |
| MOI | Mode of inheritance (AD, AR, XLD, etc.) |
Column Customization
You can show, hide, and reorder columns using the column settings menu in the toolbar. Your column preferences are saved per-user in local storage.
Sorting
Click any column header to sort by that column. Click again to reverse the sort order. Sorting is performed server-side for performance.
Row Selection
Click any row to open the Variant Details Panel. The selected row is highlighted with a blue left border.
Pagination
The table uses server-side pagination. Use the controls at the bottom of the table to navigate between pages and adjust the number of rows per page.