Variant Details
Clicking a row in the variant table opens the Variant Details Panel on the right side of the screen. This panel provides comprehensive information about the selected variant.
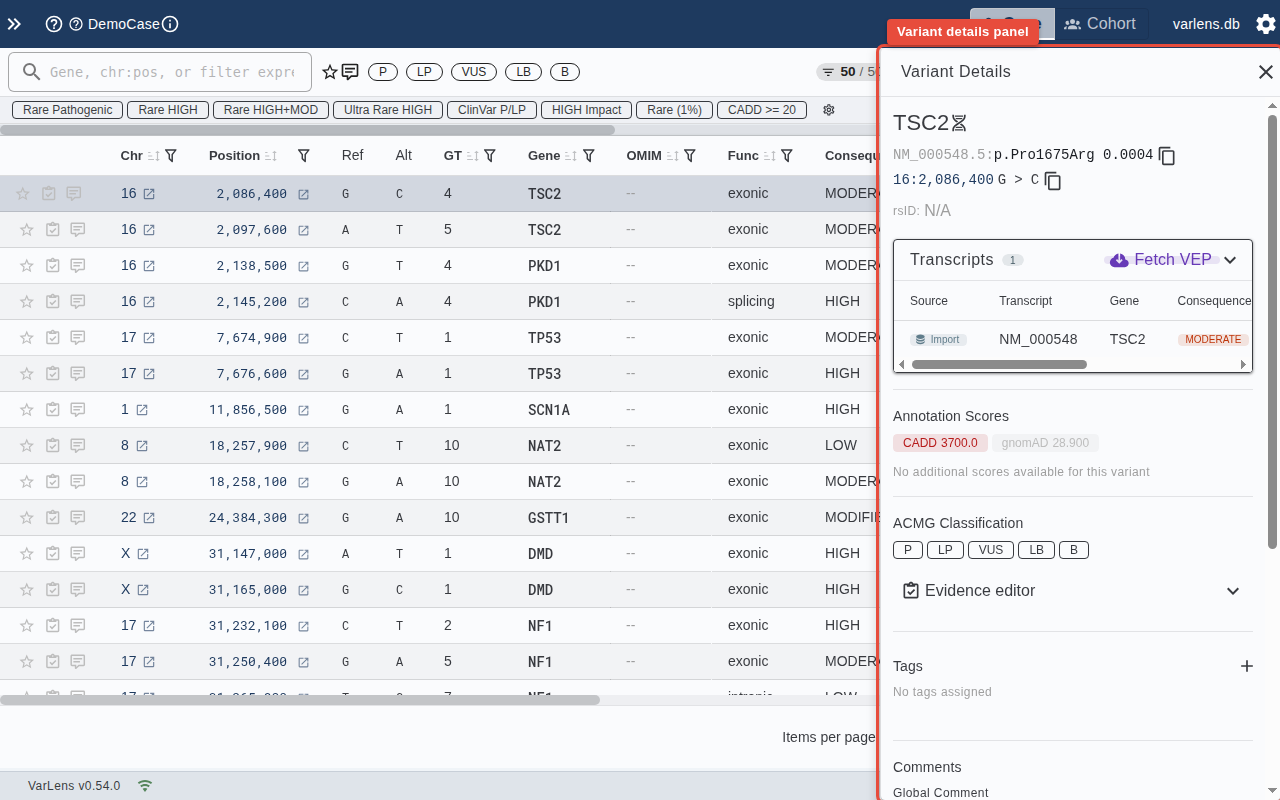
Panel Sections
Variant Identity
Shows the genomic coordinates (chr:pos:ref:alt) and any colocated variants.
Transcripts
Displays transcript annotations with MANE Select and canonical transcript indicators. You can fetch additional transcript data from VEP on demand.
Annotation Scores
Key pathogenicity and population scores:
- gnomAD AF — Population allele frequency
- CADD — Combined Annotation Dependent Depletion score
- REVEL — Rare Exome Variant Ensemble Learner score (if enriched)
- AlphaMissense — Protein structure-based pathogenicity (if enriched)
- SpliceAI — Splice prediction scores (if enriched)
ACMG Classification
Quick-classify with one-click chips (P, LP, VUS, LB, B) or open the evidence editor for detailed ACMG/AMP criteria. See Annotations for details.
Tags
Assign custom tags to organize variants for review.
Comments
Add global or per-case comments to document your analysis reasoning.
External Links
Quick links to external databases:
- UCSC Genome Browser
- gnomAD
- ClinVar
- And any custom links configured in Settings
Resizing
Drag the left edge of the panel to resize it. Your preferred width is saved.